These are notes from a recent talk where I tried to organize the landscape into three approaches: bottom-up, parallel, and uniform. Each makes a different bet about where biological structure lives and where modalities should meet inside the model.
Multimodality tasks
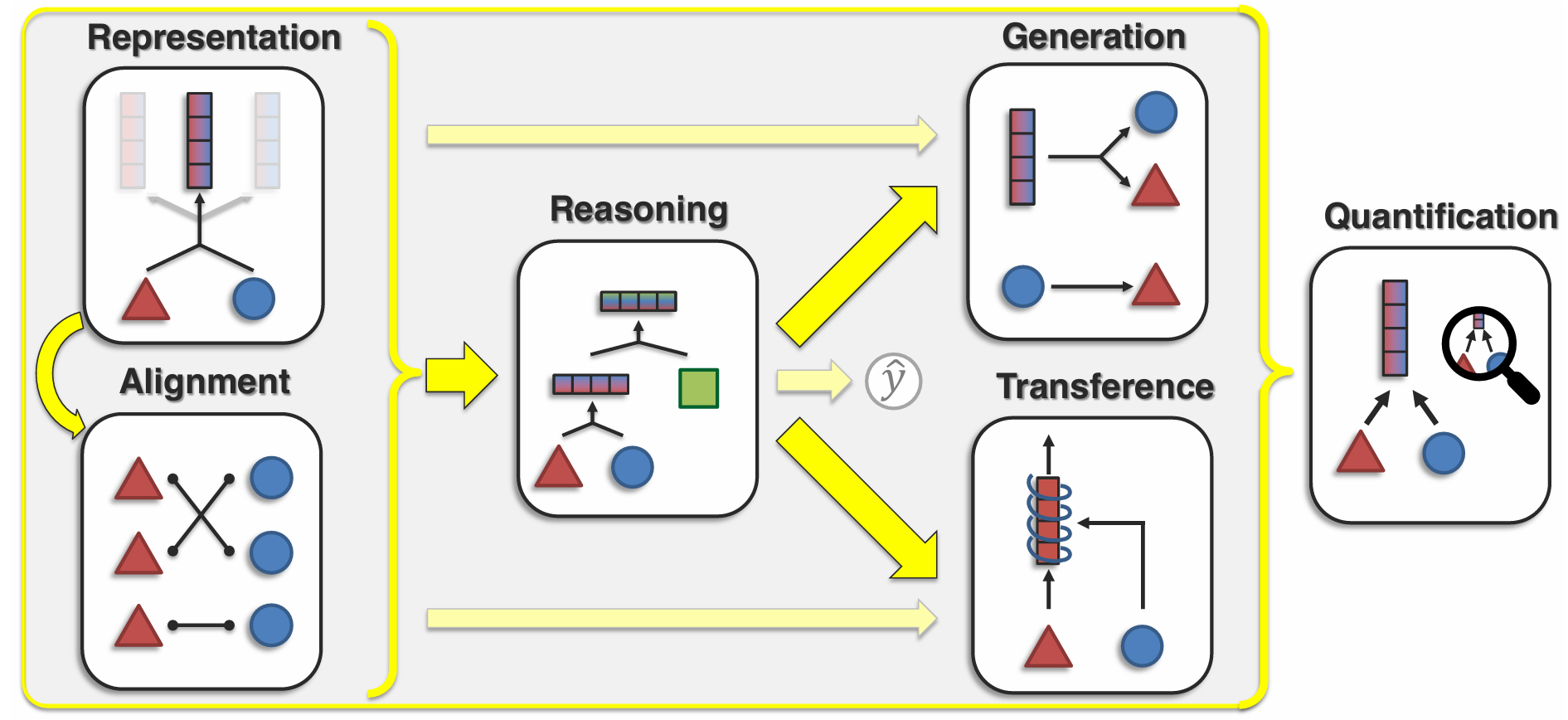
Before fixing on an architecture it helps to be explicit about the tasks we want a multimodal biological model to do — cross-modal prediction, perturbation response, cell-state inference, sequence-to-function, and so on. Different tasks pull architecture in different directions, and the rest of this post only makes sense relative to what we are asking the model to predict.
Bottom-up approach
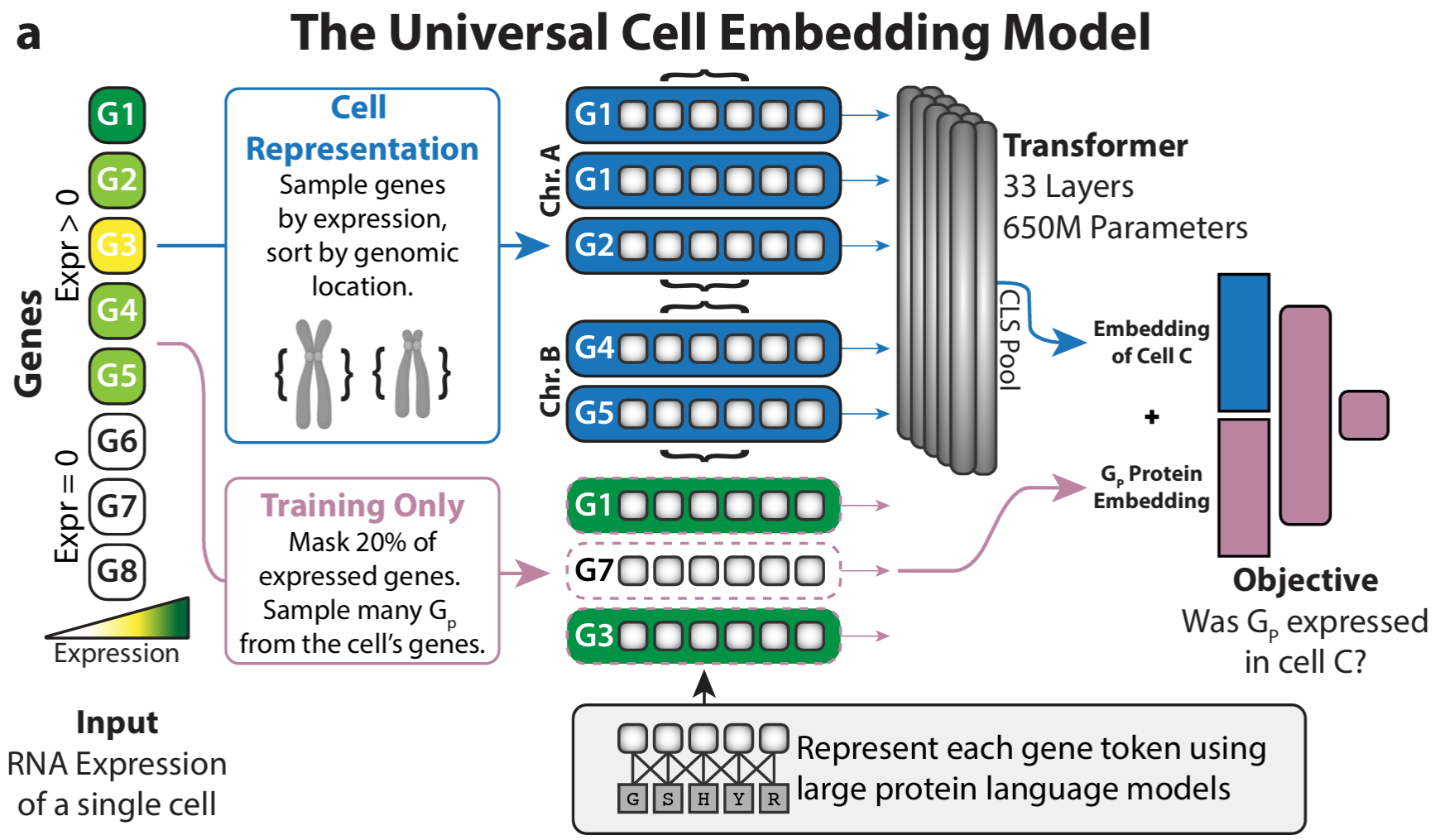
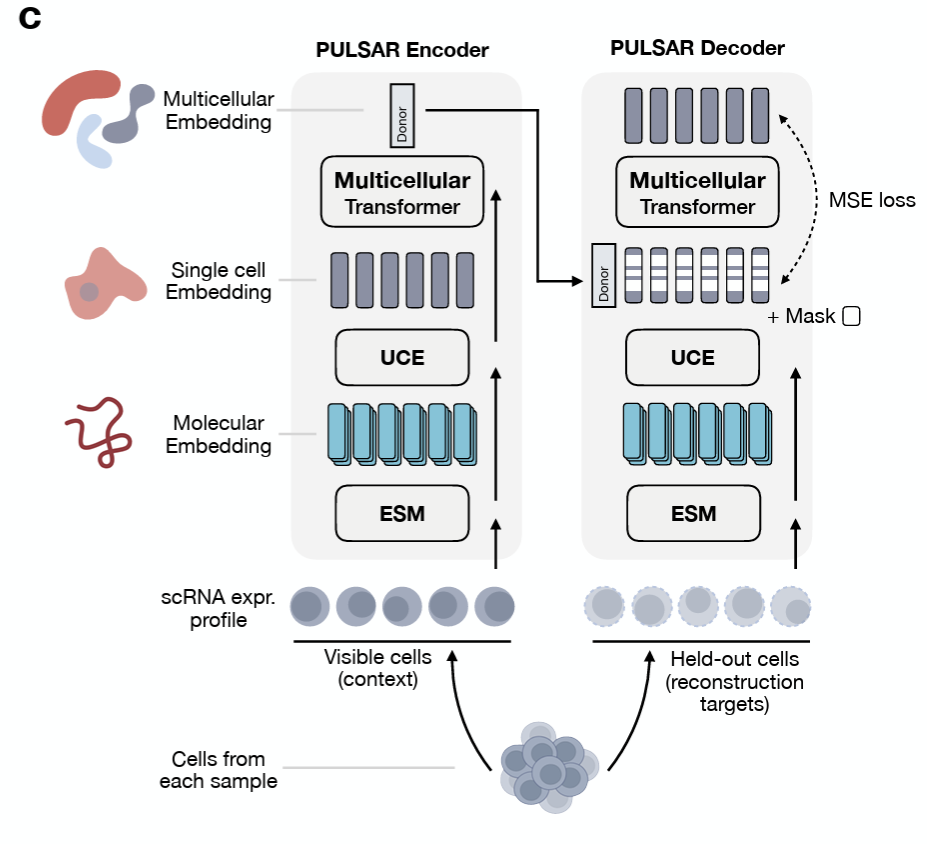
The bottom-up approach builds representations along the natural hierarchy of biology: molecular → cellular → multicellular. UCE-style models learn cell embeddings from gene-level tokens; models like PULSAR push further toward tissue- and multicellular-level structure. Each tier is trained on what is plentiful at that scale, and the next tier inherits its substrate from below.
The advantage is that each level is interpretable on its own terms and can be pretrained independently. The cost is that errors and biases compound as you climb the hierarchy.
From sequence to perturbation
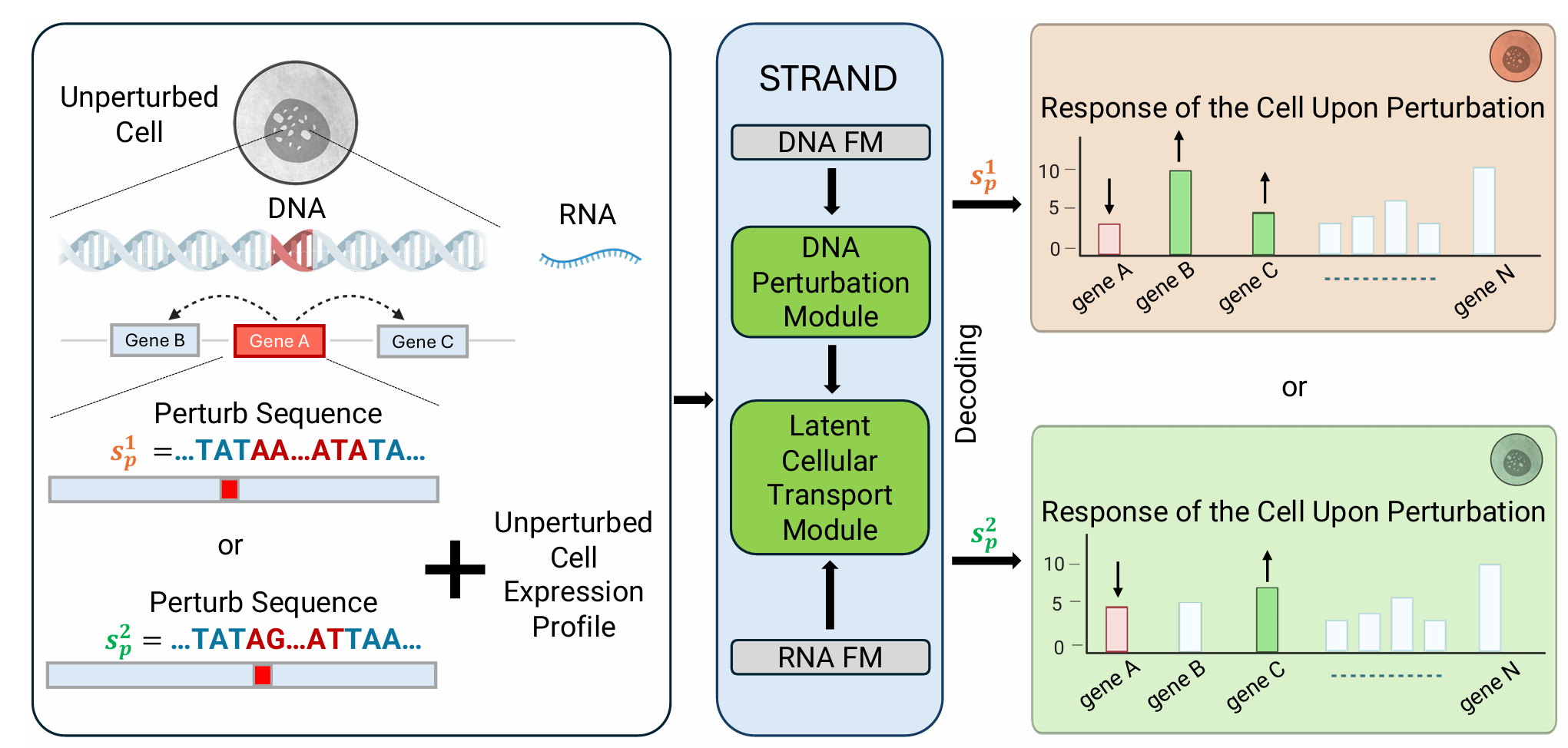
A concrete instance of the bottom-up program: start from genomic sequence and train representations that transfer downstream to perturbation prediction. The chain is sequence → expression → response, and the architectural question is at which level multimodal signals should enter.
Parallel approach
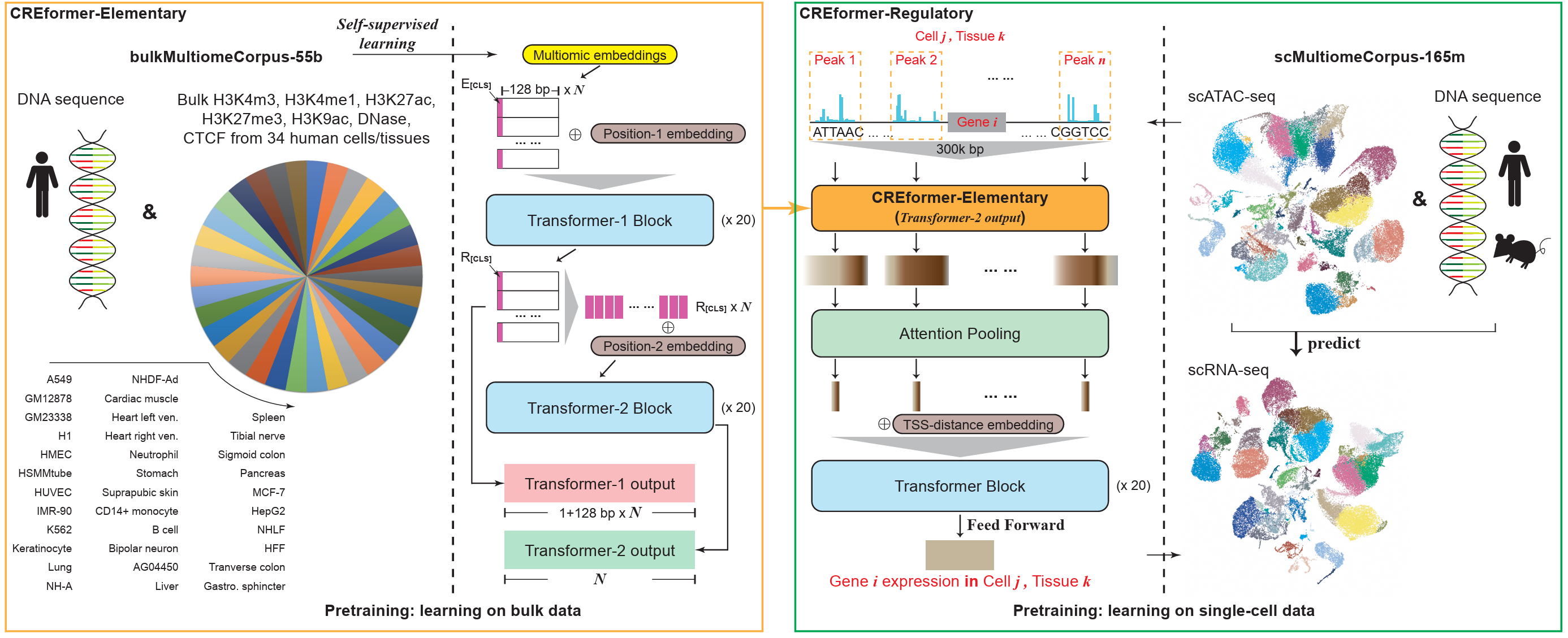
The parallel approach treats modalities as roughly co-equal and combines per-modality embeddings at the input. A canonical case: take a DNA sequence and seven epigenetic tracks, embed each independently, and directly sum the eight embeddings. Everything downstream sees a single fused vector.
This is cheap, easy to scale modality-by-modality, and trivial to extend with a new track. The price is that direct summation assumes all modalities live in the same metric space — which is rarely true biologically.
Separate encoder per modality
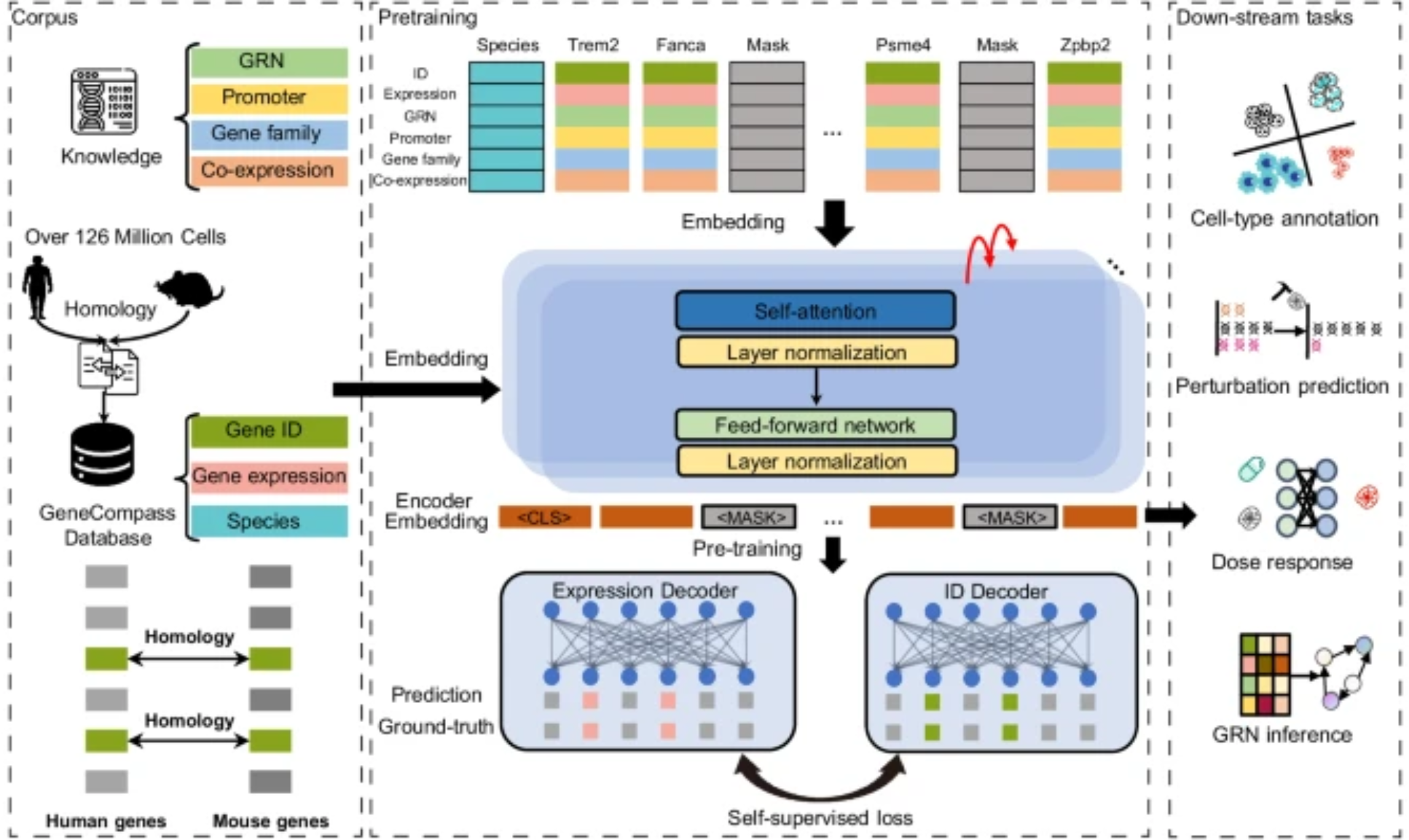
A more careful variant: keep one encoder per modality and fuse later. Each encoder can use whatever tokenization and inductive bias suits its data type, and fusion happens through concatenation, cross-attention, or gating — no longer at the input.
Different knowledge sources
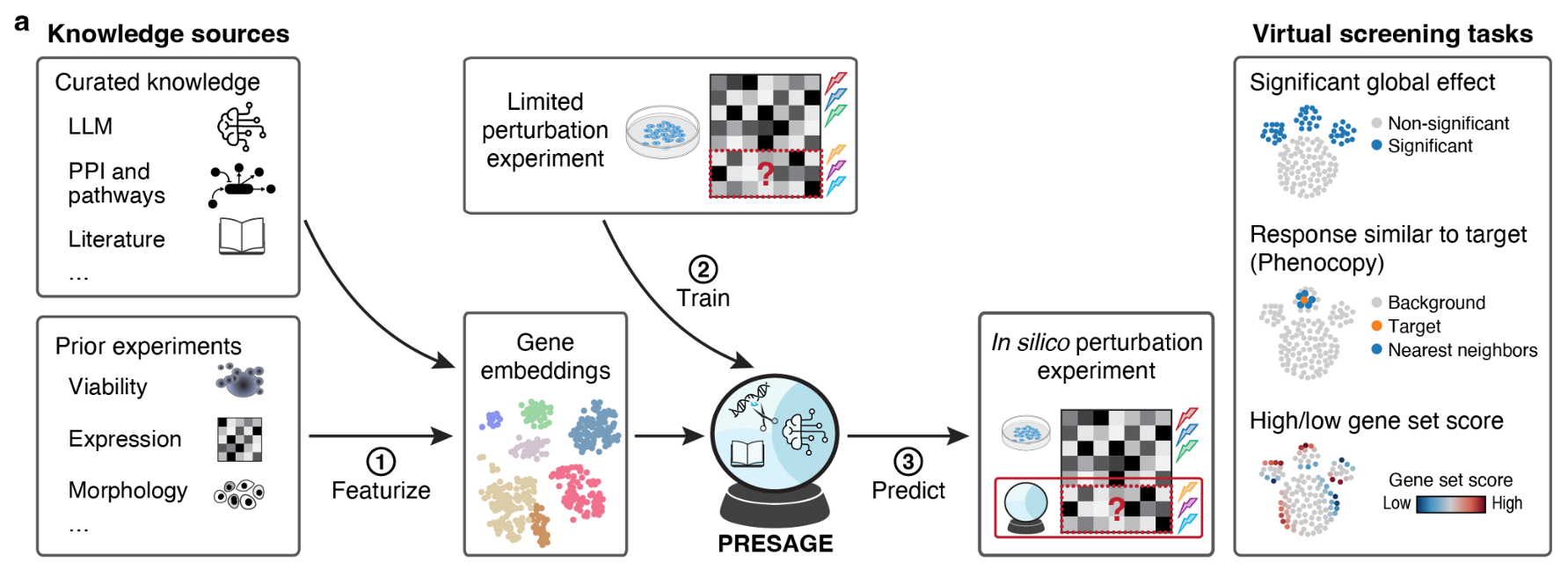
Beyond raw signals, multimodal can mean fusing different kinds of knowledge: an LLM for textual context, a knowledge graph for curated relations, tabular features for engineered priors. Two pooling strategies show up repeatedly:
- Global pooling — a weighted average of source embeddings.
- Attention-based pooling — let the query decide which source matters.
The latter usually wins when the relevance of each source varies across examples.
Uniform approach
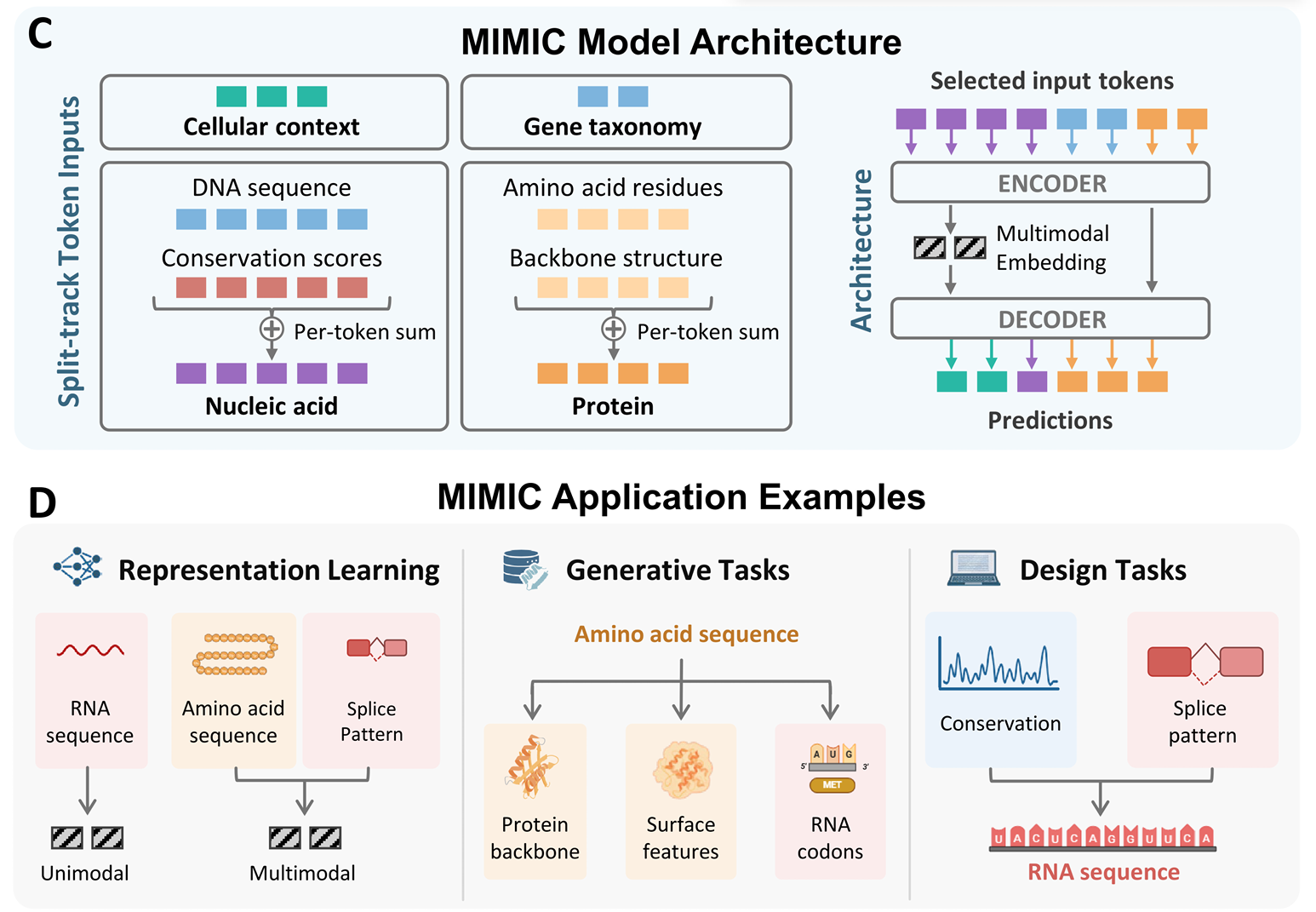
The uniform approach goes the other direction from per-modality encoders: serialize multiple sequences into a single stream and let one model digest them all. Sequence-related tasks (DNA, RNA, protein) are a natural fit — they already share a token-stream shape.
The simplicity is appealing — one model, one loss, no fusion module. The hard part is teaching a single model to respect the very different statistics of, say, codon usage versus regulatory motifs.
Relational transformer for biology
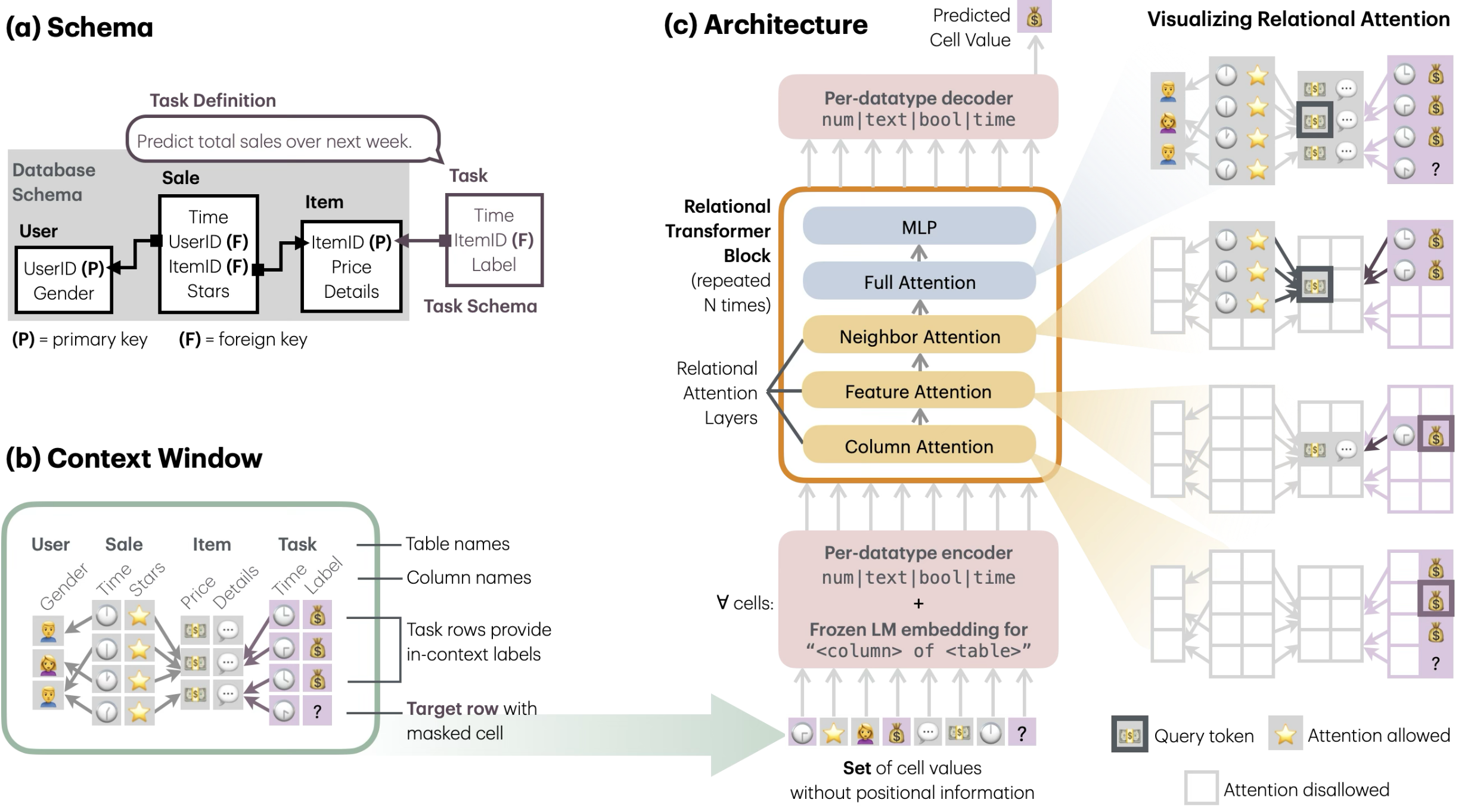
The architecture I am most interested in is a relational transformer: instead of forcing modalities through a single fusion bottleneck, represent biological entities (genes, cells, regions) as nodes and let attention range over typed relations between them.
Details
Two attention patterns carry most of the weight:
- Relational attention — for complementary modalities, where each modality contributes information the others do not. Attention selects across modalities at each layer.
- Hierarchical attention — for hierarchical modalities, where the structure itself is nested (region → gene → cell → tissue). Attention is constrained by that hierarchy.
Two open problems I keep running into:
- Memory constraint. Cross-modality attention is quadratic in token count, and biological inputs are long.
- Coupling-data constraint. Training relational attention requires examples where modalities are observed together, and truly paired multimodal datasets at scale are still rare.
These are the bottlenecks I think the next round of work — mine and others’ — needs to address.
References
- Liang, P. P., Zadeh, A., & Morency, L.-P. (2024). Foundations & Trends in Multimodal Machine Learning: Principles, Challenges, and Open Questions. ACM Computing Surveys, 56(10). https://doi.org/10.1145/3656580
- Rosen, Y., Roohani, Y., Agrawal, A., Samotorcan, L., Quake, S. R., & Leskovec, J. (2023). Universal Cell Embeddings: A Foundation Model for Cell Biology. BioRxiv. https://doi.org/10.1101/2023.11.28.568918
- Pang, K., Rosen, Y., Kedzierska, K., He, Z., Rajagopal, A., Gustafson, C. E., Huynh, G., & Leskovec, J. (2025). PULSAR: a Foundation Model for Multi-scale and Multicellular Biology. BioRxiv. https://doi.org/10.1101/2025.11.24.685470
- Fu, B., Dasoulas, G., Gabbita, S., Lin, X., Gao, S., Su, X., Ghosh, S., & Zitnik, M. (2026). STRAND: Sequence-Conditioned Transport for Single-Cell Perturbations. ArXiv Preprint ArXiv:2602.10156. https://arxiv.org/abs/2602.10156
- Yang, Z., Fan, X., Lan, M., Tang, X., Zheng, Z., Liu, B., You, Y., Tian, L., Church, G., Liu, X., & Gu, F. (2024). Multimodal foundation model predicts zero-shot functional perturbations and cell fate dynamics. BioRxiv. https://doi.org/10.1101/2024.12.19.629561
- Yang, X., Liu, G., Feng, G., Bu, D., Wang, P., & others. (2023). GeneCompass: Deciphering Universal Gene Regulatory Mechanisms with Knowledge-Informed Cross-Species Foundation Model. BioRxiv. https://doi.org/10.1101/2023.09.26.559542
- Littman, R., Levine, J., Maleki, S., Lee, Y., Ermakov, V., Qiu, L., Wu, A., Huang, K., Lopez, R., Scalia, G., Biancalani, T., Richmond, D., Regev, A., & Hütter, J.-C. (2025). Gene-embedding-based prediction and functional evaluation of perturbation expression responses with PRESAGE. BioRxiv. https://doi.org/10.1101/2025.06.03.657653
- Golkar, S., Kovalic, J., Espejo Morales, I., Sledzieski, S., Cho, K., Cranmer, M., Ho, S., & others. (2026). MIMIC: A Generative Multimodal Foundation Model for Biomolecules. ArXiv Preprint ArXiv:2604.24506. https://arxiv.org/abs/2604.24506
- Ranjan, R., Hudovernik, V., Znidar, M., Kanatsoulis, C., Upendra, R., Mohammadi, M., Meyer, J., Palczewski, T., Guestrin, C., & Leskovec, J. (2025). Relational Transformer: Toward Zero-Shot Foundation Models for Relational Data. ArXiv Preprint ArXiv:2510.06377. https://arxiv.org/abs/2510.06377






